Actualités
-

-
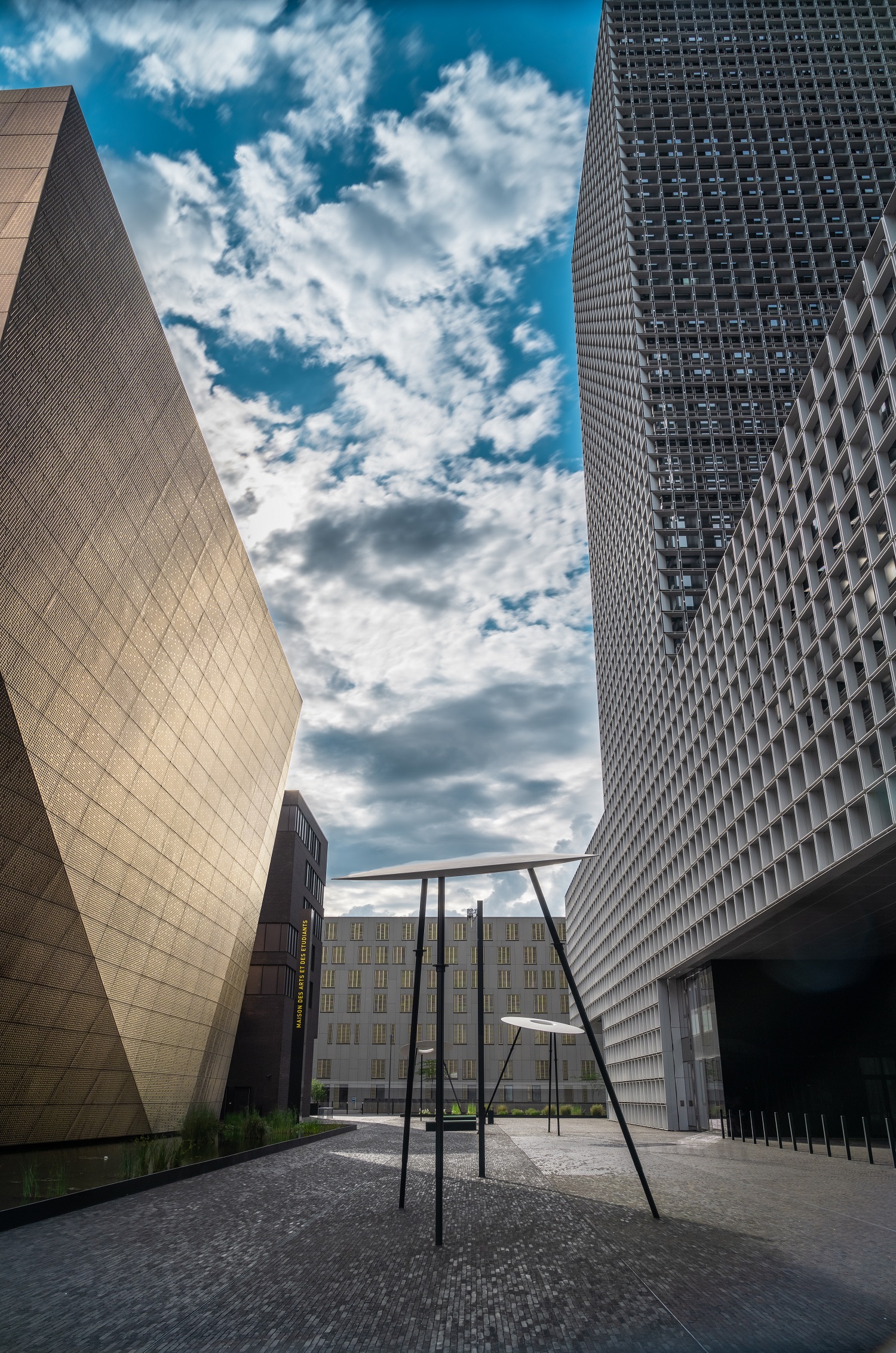

Un plan d’action pour faire progresser l’évaluation de la recherche
Recherche, UniversitéEn savoir plus -

-
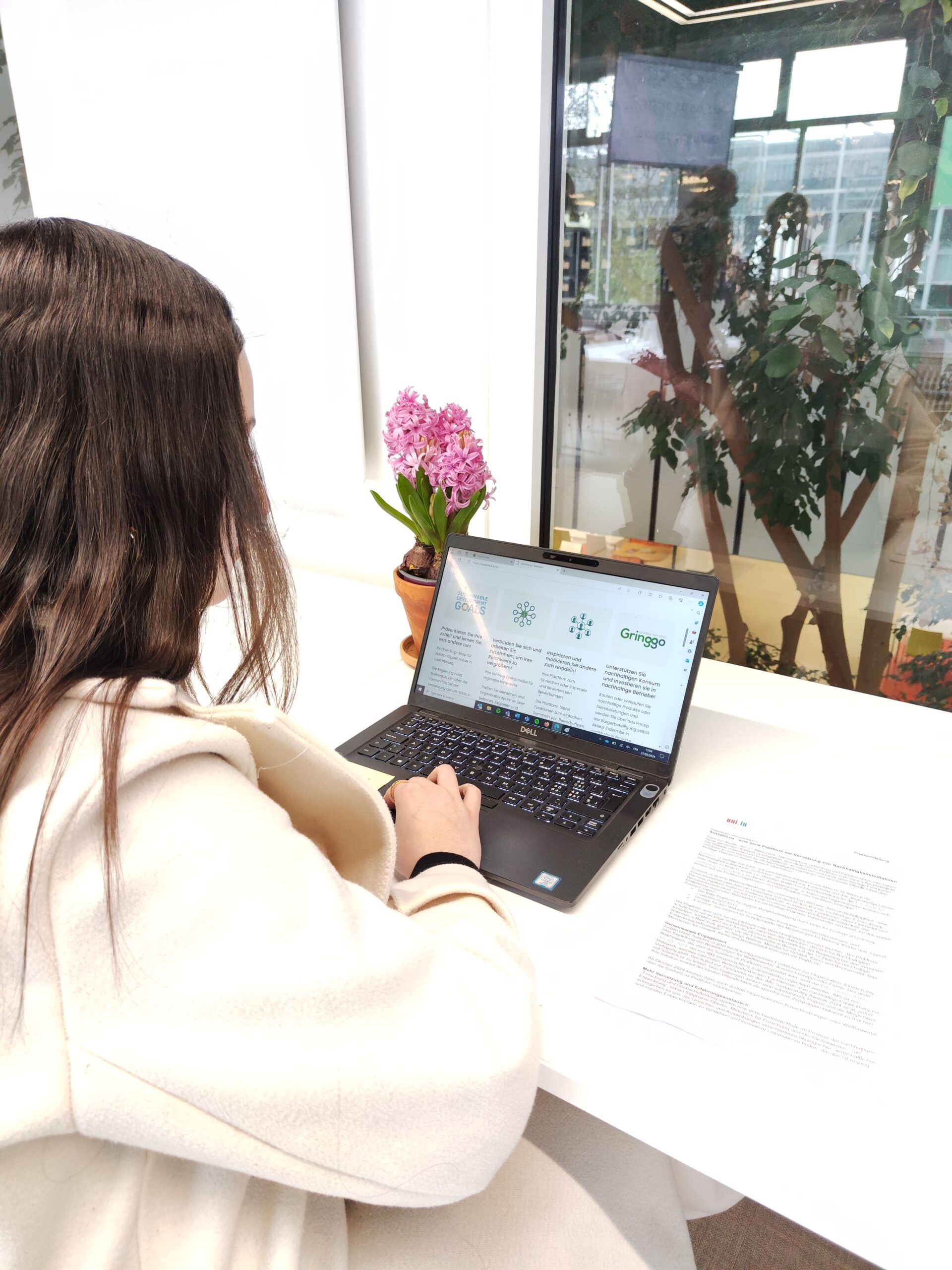

Avec SustainLux, le développement durable est à portée de clic
Relations avec le publicDurabilité, Sciences socialesEn savoir plus -
-
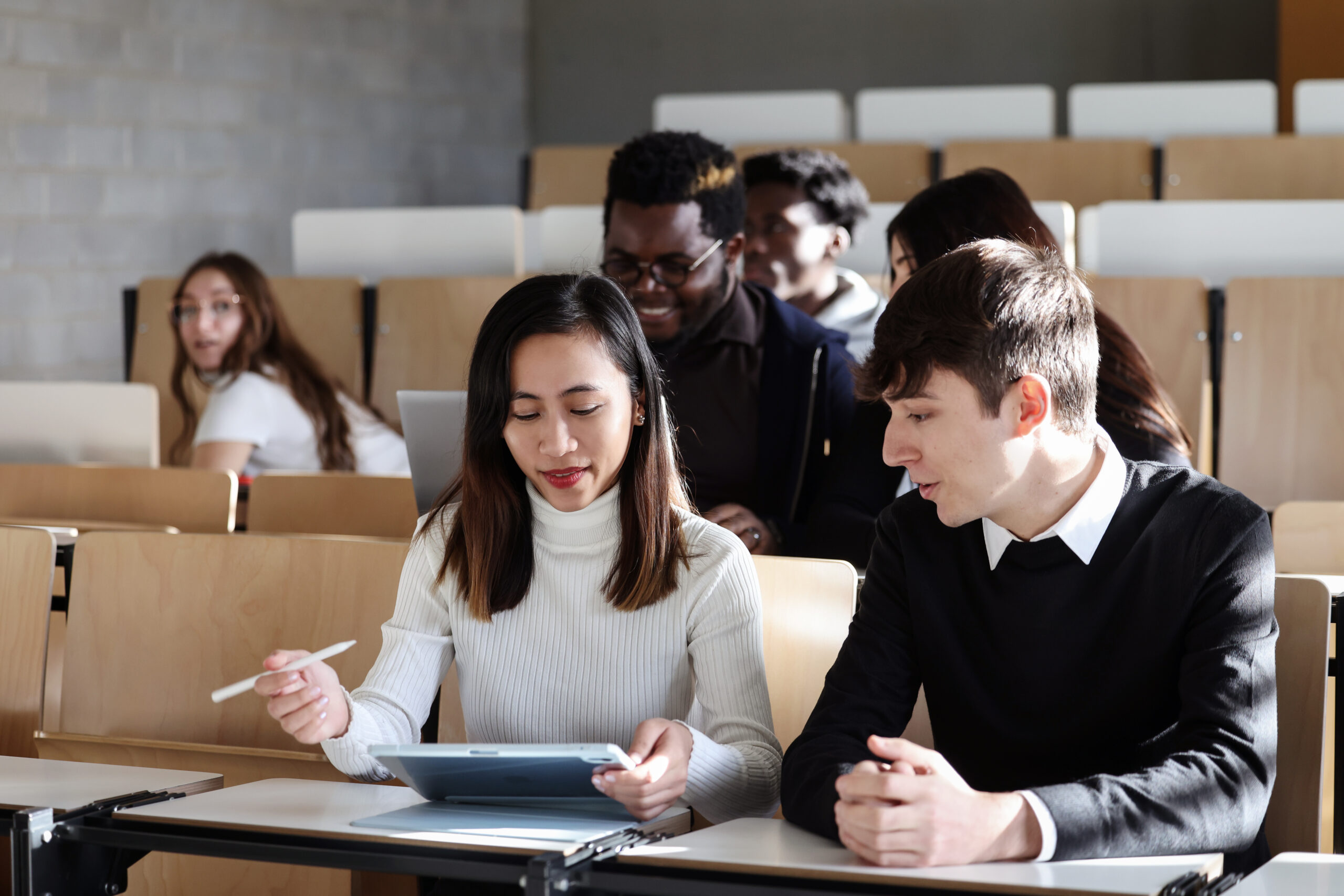

Du neuf dans l’offre d’études pour la rentrée 2024-2025
Communiqués de presse, Education, UniversitéDroit, Informatique & TIC, Ingénierie, Sciences de la vie & médecine, Sciences humainesEn savoir plus
Appel à candidatures
-

-

Appel à candidatures : Directeur fondateur /Directrice fondatrice du Centre Interdisciplinaire en Droit Européen
UniversitéEn savoir plus