Actualités
-

-

-

Découvrir la communication scientifique grâce à nouveau livre en libre accès
Communiqués de presse, Relations avec le public, UniversitéEn savoir plus -

Un plan d’action pour faire progresser l’évaluation de la recherche
Recherche, UniversitéEn savoir plus -
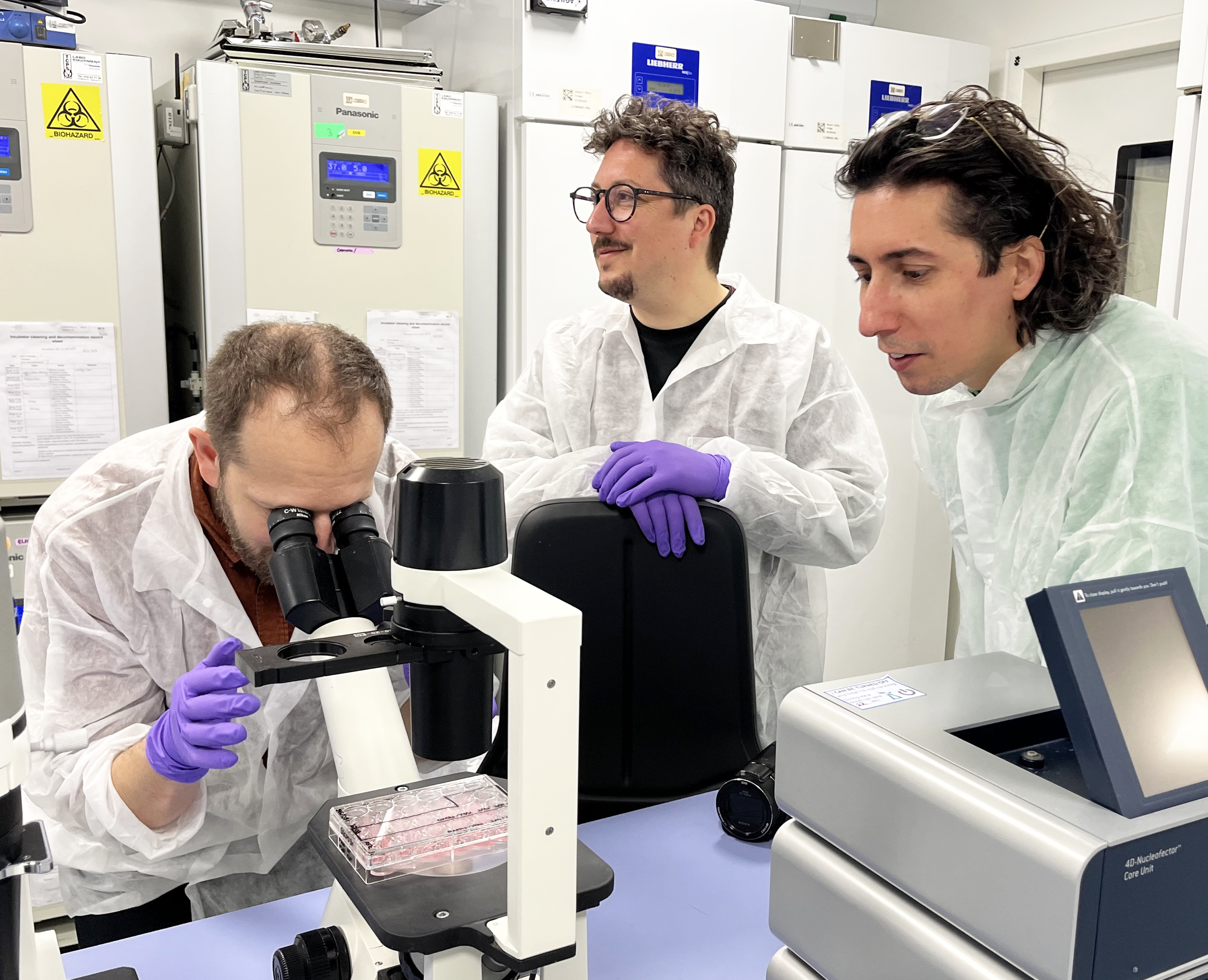

L’art à la rencontre de la science : Les premiers artistes en résidence à l’Université
La vie sur le campus, Relations avec le publicSciences de la vie & médecineEn savoir plus -

Appel à candidatures
-

-

Appel à candidatures : Directeur fondateur /Directrice fondatrice du Centre Interdisciplinaire en Droit Européen
UniversitéEn savoir plus